 |

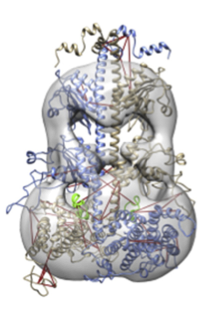
IMP's broad goal is to contribute to a comprehensive structural characterization of biomolecules ranging in size and complexity from small peptides to large macromolecular assemblies, by integrating data from diverse biochemical and biophysical experiments. IMP provides an open source C++ and Python toolbox for solving complex modeling problems, and a number of applications for tackling some common problems in a user-friendly way. IMP can also be used from the Chimera molecular modeling system, or via one of several web applications.
IMP is open source software, mostly available under the terms of the GNU Lesser General Public License (LGPL). (Some IMP modules are available under the GNU GPL instead.)
Please note: While IMP currently supports both Python 2 and Python 3, Python 2 was officially retired several years ago, and we plan to drop support for Python 2 in the next IMP stable release after June 30th, 2024. If this would cause a lot of issues with your own workflows and for some reason you cannot move to Python 3, please let us know or update the relevant GitHub issue.
Get started with IMP by downloading it and checking out the documentation.

The IMP software is used as part of the National Center for Dynamic Interactome Research (NCDIR).
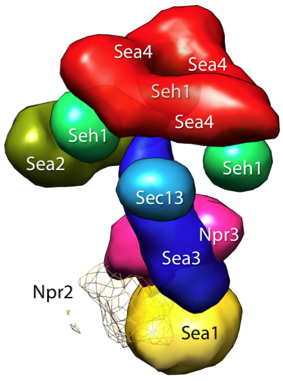
If you use IMP, please cite D. Russel, K. Lasker, B. Webb, D. Schneidman, J. Velázquez-Muriel, A. Sali, "Putting the pieces together: integrative structure determination of macromolecular assemblies", PLoS Biology, 2012. The main page of each IMP module in the documentation also lists publications relevant to that module.