These scripts demonstrate the use of IMP, MODELLER, and PMI in the modeling of the solution structure of the Nef-CD4-AP2 complex using 90 DSSO chemical cross-links and the X-ray structure of the crystal structure, a comparative model of the β2 subunit 24-89 region built based on the AP2 structure.
First, MODELLER is used to generate comparative model of the β2 subunit 24-89 region. Then, IMP is used to model these components using crystal structure and the DSSO crosslinks to obtain the solution structure of the Nef-CD4(CD)-AP2(Δμ2-CTD) complex.
The modeling protocol will work with a default build of IMP, but for most effective sampling, IMP should be built with MPI so that replica exchange can be used.
-
datacontains all relevant data, input structures that were generated by MODELLER or deposited in PDB, etc.mod_extended_AP2beta2.pyMODELLER script to add the β2 subunit 24-89 region built based on the AP2 structure using MODELLER
-
scriptsmod_Nef_AP2_newhelix.pythe main IMP/PMI script modeling with 3 crystal interfaces
-
scripts/analysisThe best 500 models from the modeling are accumulated in this directory, and updated as the calculation proceeds. -
scripts/figuresThe structures of the lowest temperature replica will be written here as RMF files. -
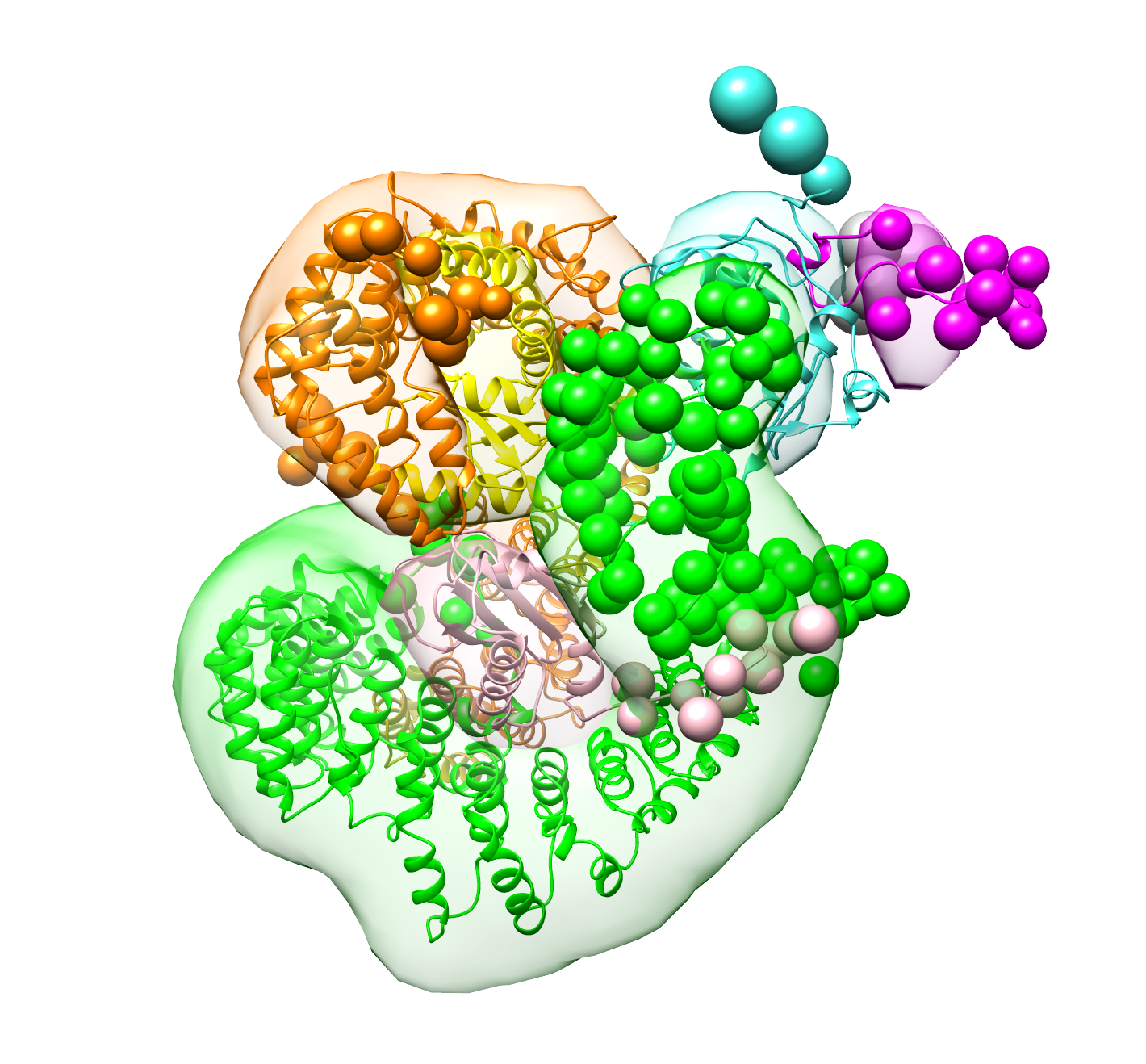
outputsFor reference, the models described in the Nup84 publication are deposited in this directory. For each of the two clusters discovered in the study, the cluster representative (the best scoring individual model in the cluster) is available in RMF format, together with the top five best scoring models in PDB format. An accompanying stat file contains useful statistics on the simulation, such as whether each of the crosslinks was satisfied. The localization densities, as shown in Figure 6 on the publication are also available, as Chimera session files.
cd data/(python mod_extended_AP2beta2.py > mod_extended_AP2beta2.log)
cd scriptspython mod_Nef_AP2_newhelix.py &> mod_Nef_AP2_newhelix.out(on a single processor; prependmpirun -np 4or similar if you built IMP with MPI support)- You can add the
--testoption to the command line to quickly test the script; this simply runs the sampling for fewer steps (should complete in an hour or two).
Next, analyze resulting models (this script can also be used to combine results from multiple independent runs):
cd scripts/analysispython run_analysis.py ../- (If you used the
--testoption above, use it here too.)
Author(s): Ignacia Echeverria
Date: May 5th, 2020
License: LGPL. This library is free software; you can redistribute it and/or modify it under the terms of the GNU Lesser General Public License as published by the Free Software Foundation; either version 2 of the License, or (at your option) any later version.
Testable: Yes.
Parallelizeable: Yes
Publications:
- Kwon, Y., Kaake, R.M., Echeverria, I. et al. Structural basis of CD4 downregulation by HIV-1 Nef. Nat Struct Mol Biol (2020). (https://doi.org/10.1038/s41594-020-0463-z).



 Download files
Download files Verified to work with the
Verified to work with the  To install the software needed to reproduce this system with the
To install the software needed to reproduce this system with the
 To set up the environment on the UCSF Wynton cluster to run
this system, run:
To set up the environment on the UCSF Wynton cluster to run
this system, run: