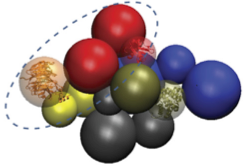
Data and scripts to model eIF3 from MS data.
The method used here is the same as that used for the MS benchmark.
To generate models, run the ms_cg.py script. This will generate a number of models of the complex, named conf_eif3.<num>.pym.
The .pym files can be converted to .tcl files by running the pym2tcl.py script, and then to .mfj files by running the tcl2mfj.py script.
Author(s): Argyris Politis, email:argyris.politis@kcl.ac.uk
Version: 1.0
License: LGPL. This library is free software; you can redistribute it and/or modify it under the terms of the GNU Lesser General Public License as published by the Free Software Foundation; either version 2 of the License, or (at your option) any later version.
Testable: Yes.
Parallelizeable: No
Publications:
- Keren Lasker, Javier A. Velazquez-Muriel, Benjamin M. Webb, Zheng Yang, Thomas E. Ferrin, Andrej Sali, Macromolecular assembly structures by comparative modeling and electron microscopy, Methods in Molecular Biology, 2011.
- Politis A, Stengel F, Hall Z, Hernandez H, Leitner A, Waltzhoeni T, Robinson CV, Aebersold R. A mass spectrometry-based hybrid method for structural modelling of protein complexes, Nature Methods, 11, 403-406, (2014)
- A. Politis, C. Schmidt, E. Tjioe, A. Sandercock, K. Lasker, Y. Gordiyenko, D. Russel, A. Sali, C. Robinson. Topological models of heteromeric protein assemblies from mass spectrometry: Application to the yeast eIF3:eIF5 complex, Chem Biol 22, 117-128, 2015.



 Download files
Download files Verified to work with the
Verified to work with the  To install the software needed to reproduce this system with the
To install the software needed to reproduce this system with the
 To set up the environment on the UCSF Wynton cluster to run
this system, run:
To set up the environment on the UCSF Wynton cluster to run
this system, run: