This repository contains all the scripts for the sampling exhaustiveness tests along with the inputs for the benchmark cases and results on the benchmarks.
The inputs folder contains inputs for sampling for the benchmark cases,such as PDB and crosslink files.
The scripts folder contains scripts for sampling IMP models and scoring ZDOCK models, as well as the scripts for the protocol and plotting outputs from the protocol.
The results folder contains the models sampled by IMP, and the resulting clusters of good-scoring models, along with the top models from ZDOCK and the outputs of the sampling exhaustiveness protocol.
Please refer to the README.md file in each directory for specific details.
Author(s): Shruthi Viswanath, Ilan E. Chemmama
Maintainer: shruthivis,ichem001
License: LGPL This library is free software; you can redistribute it and/or modify it under the terms of the GNU Lesser General Public License as published by the Free Software Foundation; either version 2 of the License, or (at your option) any later version.
Publications:
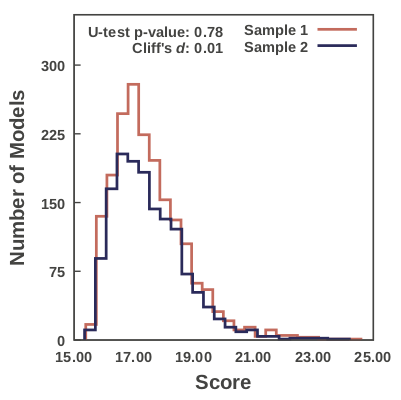
- S. Viswanath, I.E.Chemmama, P. Cimmermancic and A. Sali, Validating Exhaustiveness of Stochastic Sampling for Integrative Modeling of Macromolecular Structures, Biophysical Journal 113, 2344-2353, 2017.


 Download files
Download files Warning: these files have not yet been verified to work with the latest version of IMP. We will update this page when they have been.
The files are also available at
Warning: these files have not yet been verified to work with the latest version of IMP. We will update this page when they have been.
The files are also available at  To install the software needed to reproduce this system with the
To install the software needed to reproduce this system with the
 To set up the environment on the UCSF Wynton cluster to run
this system, run:
To set up the environment on the UCSF Wynton cluster to run
this system, run: