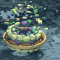News
Postdoctoral positions at the Quantitative Biology Institute available:
The Quantitative Biology Institute (QBI) at the University of California, San Francisco, is recruiting six postdoctoral fellows in the area of computational structural biology. See the description of these positions for more information.
PDB creates an archive for integrative structures:
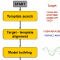
wwPDB has created a new archive, PDB-Dev,
for storing structural models obtained through use of integrative methods.
It contains a rapidly-growing number of such models, produced with software
including our own IMP package.
Congratulations to Daniel:
Daniel Saltzberg has received the Lilly Research Award!
Congratulations to SJ:
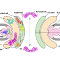
The paper "Integrative Structure and Functional Anatomy of a Nuclear Pore Complex" has been published in Nature!
Congratulations to Sara:
The paper "Prediction of enzymatic pathways by integrative pathway mapping" has been published in eLife!
Congratulations to Sara:
Sara Calhoun received the Mel Jones Excellence in Graduate Student Research Award !
Congratulations to Adrian:
Adrian Stecula is a recipient of an AFPE Pre-Doctoral Award in Pharmaceutical Science!
Congratulations to Ilan:
Ilan Chemmama has received a 3 year NSF fellowship !
Congratulations to Kate:
Kate Stafford has received a 3 year NRSA fellowship !
Congratulations to Adrian:
Adrian is a recipient of the PhRMA fellowship in Pharmacology/Toxicology !
Congratulations to Natalia:
Natalia is a recipient of a 2013-2014 Merit Fellowship Award - an Achievement Rewards for College Scientists (ARCS) Scholarship !
Congratulations to SJ:
The paper "Structure, Dynamics, Evolution, and Function of a Major Scaffold Component
in the Nuclear Pore Complex", P. Sampathkumar, et al, has been published in Structure!
We also congratulate Graham Johnson of UCSF for the great cover image!
Congratulations to Javi:
We wish him the best in his position as Bioinformatics Analyst at Cypher Genomics!
Congratulations to Avner:
We wish him the best in his new position as Assistant Professor at Mount Sinai Hospital !
Congratulations to Jeremy and Peter:
Jeremy won the Frank M. Goyan Award for Excellence in Physical Chemistry and Peter has received the 2012 School of Pharmacy Research Award !
Congratulations to Keren:
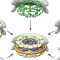
The paper "Molecular architecture of the 26S proteasome holocomplex determined by an integrative approach", K. Lasker, et al, has been accepted for publication in PNAS !
Congratulations to Riccardo on the news of his Swiss National Science Foundation fellowship!!
Congratulations to Avner:
The paper "Structure-based discovery of prescription drugs that interact with the norepinephrine transporter, NET", A. Schlessinger, E. Geier, H. Fan, J. Irwin, B. Shoichet, K. Giacomini, A. Sali, has been accepted for publication in PNAS !
Congratulations to Peter on being awarded the Howard Hughes Medical Institutes Predoctoral Fellowship!!
Congratulations Charles on being awarded the NSF Graduate Research Fellowship!!
Selected Publications
-
S. Otsuka, J.O.B. Tempkin, W. Zhang, A.Z. Politi, A. Rybina, M.J. Hossain, M. Kueblbeck, A. Callegari, B. Koch, N.R. Morero, A. Sali, J. Ellenberg.
"A quantitative map of nuclear pore assembly reveals two distinct mechanisms" Nature 613, 575-581, 2023.


-
B. Raveh, L. Sun, K.L. White, T. Sanyal, J. Tempkin, D. Zheng, K.B. Pilla, J. Singla, C. Wang, J. Zha, A. Li, N.A. Graham, C. Kesselman, R.C. Stevens, A. Sali.
"Bayesian metamodeling of complex biological systems across varying representations" Proc Natl Acad Sci USA 118, e2104559118, 2021.


-
S.J. Kim, J. Fernandez-Martinez, I. Nudelman, Y. Shi, W. Zhang, B. Raveh, T. Herricks, B.D. Slaughter, J. Hogan, P. Upla, I.E. Chemmama, R. Pellarin, I. Echeverria, M. Shivaraju, A.S. Chaudhury, J. Wang, R. Williams, J.R. Unruh, C.H. Greenberg, E.Y. Jacobs, Z. Yu, M.J. de la Cruz, R. Mironska, D.L. Stokes, J.D. Aitchison, M.F. Jarrold, J.L. Gerton, S.J. Ludtke, C.W. Akey, B.T. Chait, A. Sali, M.P. Rout. "Integrative Structure and Functional Anatomy of a Nuclear Pore Complex" Nature 555, 475-482, 2018.


-
A. Sali. "From integrative structural biology to cell biology" J Biol Chem 100743, 2021.


This is not an official UCSF website. The opinions or statements expressed herein should not be taken as a position of or endorsement by the University of California, San Francisco.