|
|
Frequently asked questions (FAQ) and examples
Please also check the modeller_usage
mailing list archives where your question may have already been answered.
Modeller Installation
Modeller Usage 001
Modeller Usage
- I do not care about the details of a model, I only want to calculate
it very fast to get a quick idea about how it looks or to confirm that
my alignment is clearly unreasonable in the structural sense.
- How can I refine the model in successive steps?
- I want to model one or more loops very thoroughly (meaning spending
a lot of CPU time, not necessarily modeling more accurately).
- I want to build a model of a chimeric protein based on two
known structures. Alternatively, I want to build a multi-domain protein
model using templates corresponding only to the individual domains.
- I don't want to use one region of a template for construction
of my model.
- I want to define (additional) disulfide bonds in the target
sequence because no equivalent disulfide bonds exist in any of the
templates (in which case PATCH_SS_TEMPLATES cannot define them
automatically).
- I want to explicitly force certain Pro residues to the
cis
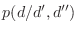 conformation. conformation.
- How can I select/remove/add a set of restraints?
- I want to add my own restraints for optimization of the model.
- I want to add my own restraints to the file with the
automatically derived homology restraints, immediately after the
default calculation of the homology-derived restraints.
- I have my own restraints file to be used exclusively
for optimization by the default comparative modeling routine.
- I have my own initial structure to be used for optimization
by the default comparative modeling routine.
- What are the different refinement levels really doing?
- I want to change the default optimization schedule.
- I want to build an all hydrogen atom model with water molecules and
other non-protein atoms (atoms in the HETATM records in the PDB file).
- How do I build a model with water molecules or residues that
do not have an entry in the topology and/or parameter files?
- How do I define my own residue types, such as D-amino acids,
special ligands, and unnatural amino-acids?
- How do I define my own patching residue types?
- Is it possible to restrain secondary structure in the
target sequence?
- I want to patch the N-terminal or (C-terminal) residue (e.g.,
to model acetylation properly), but the PATCH command does not work.
- Is it possible to use templates with the coordinates for
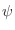 atoms only? atoms only?
- How do I analyze the output log file?
- How do I prevent ``knots'' in the final models?
- What do I do when I get Syntax error at line 1: `(' unexpected message?
- What is considered to be the minimum length of a sequence motif
necessary to derive meaningful constraints from the alignment to use in
modeling.. one, two, three, or more?
- Does Modeller have a graphical interface (GUI) ?
Modeller Installation
- Installation under Windows
MODELLER 6v2 for MS Windows INSTALLATION.
1. Uncompress the mod6v2.zip package into your C: drive (all apropriate
directories will be created under c:\modeller6v2 directory). You may want to
put the files into another directory and/or drive, in this case simply
uncompress then to other drive and/or move the main directory (modeller6v2)
together with all subdirectories and files wherever you want.
2. There are two files with environmental variables for MODELLER:
c:\modeller6v2\bin\mod.bat and c:\modeller6v2\bin\modenv.bat .
You need to edit these files and replace XXX in the line
set KEY_MODELLER6v2=XXX
with an appropriate MODELLER key (see MODELLER documentation for more
details). You also need to change the line
set MODINSTALL6v2=c:\mod6v2 to
set MODINSTALL6v2=c:\modeller6v2
(path has been changed)
3. If you have placed your MODELLER in different drive and/or directory
you need to replace c:\modeller6v2 in the line
set MODINSTALL6v2=c:\modeller6v2
with your drive and/or directory.
4. In order to run MODELLER from any script you need to add the following line
in your script file before calling modeller:
call c:\modeller6v2\bin\modenv.bat
5. You may execute modeller in a windows command prompt by calling
c:\modeller6v2\bin\mod.bat [arguments]
6. You may want to add the line call c:\modeller6v2\bin\modenv.bat
to your configuration script.
7. In order to use compressed PDB files from MODELLER a MS Windows version of
GNU gzip (available on the Internet) has to be installed and placed in
the search path.
- Modeller execution under Windows from the command line
Modeller does not have a graphical user interface but is a command line tool.
You have to open a command prompt window and execute modeller with giving it an argument:
Programs-Accessories-CommandPrompt gives you the command prompt window
The Readme file states:
5. You may execute modeller in a windows command prompt by calling
c:\modeller6v2\bin\mod.bat [arguments]
So, first I find the complete path of my modeller installation.
Please be sure not to have any spaces in directory names, modeller cannot handle that.
My modeller installation is in C:\modeller6v2
Then, I check my mod.bat files and modenv.bat files:
modenv.bat:
set MODINSTALL6v2=c:\modeller6v2
set KEY_MODELLER6v2=XXX
set LIBS_LIB6v2=%MODINSTALL6v2%\modlib\libs.lib
set EXECUTABLE_TYPE6v2=i386-windf
set PATH=%MODINSTALL6v2%\bin;%PATH%
mod.bat:
set MODINSTALL6v2=c:\modeller6v2
set KEY_MODELLER6v2=XXX
set LIBS_LIB6v2=%MODINSTALL6v2%\modlib\libs.lib
set EXECUTABLE_TYPE6v2=i386-windf
set PATH=%PATH%;%MODINSTALL6v2%\bin
%MODINSTALL6v2%\bin\modeller6v2 %1 %2 %3 %4 %5 %6 %7 %8 %9
Then I executed modeller:
C:\modeller6v2\examples\tutorial-model>c:\modeller6v2\bin\mod.bat model-default.top
(all one line)
- Modeller execution of run_tops1 under Windows
The run_tops1 file is not a modeller top file. It is a script that runs a
number of modeller jobs. This is the content:
run_tops1:
#!/bin/sh
mod6v2 initial
mod6v2 model-default
mod6v2 model-segment
mod6v2 model-fast
mod6v2 model-loop
# procheck 1fdx.B999901 2.0
mod6v2 compare
It is a unix shell script.
In a unix environment, you would execute it like this:
cd whereever/examples/tutorial-model
./run_tops1
all those top files (initial.top, model-default.top, etc.) are in this
directory.
For windows, you should remove the first line of the script and the procheck
line (commented out in the unix script), and rename it run_tops1.bat. Then
you should change mod6v2 to mod, execute the modenv.bat file to set the
environment, and execute it as run_tops1 from the tutorial-model directory.
Modeller Usage 001
- How do I run modeller successfully?
If you are getting an error message similar to the following:
fullfn__230E> File not found: mod
Directories: :${MODINSTALL6v2}/bin/
Extensions : .top:
Prefixes : nothing, pdb
chances are that you didn't give modeller an argument.
You should try the following:
change directory into the examples/tutorial-model directory
run the following command (assuming that the modeller executable is in your path):
"mod initial"
> exchange mod with your executable, e.g. mod6v2, or leave mod for windows, initial is
> refering to the initial.top file in the current directory.
> Then execute ls -l (or dir for windows) in this directory to identify the output files.
Modeller Usage
- I do not care about the details of a model, I only want to calculate
it very fast to get a quick idea about how it looks or to confirm that
my alignment is clearly unreasonable in the structural sense.
Only one model can be calculated by this routine because the
starting structure is not randomized before optimization. Only
a very limited amount of the variable target function optimization
with conjugate gradients is done. This is usually for a factor of 3
faster than the default procedure. For example, it takes about
17 seconds of CPU time to model a 60-residue protein on an SGI
workstation with a R10000-195 processor.
# Very fast homology modelling by the MODELLER TOP routine 'model'.
INCLUDE # Include the predefined TOP routines
SET ALNFILE = 'alignment.ali' # alignment filename
SET KNOWNS = '5fd1' # codes of the templates
SET SEQUENCE = '1fdx' # code of the target
SET ATOM_FILES_DIRECTORY = './:../atom_files' # directories for input atom files
SET STARTING_MODEL = 2
SET ENDING_MODEL = 2
SET OUTPUT_CONTROL = 1 1 1 1 1
# SET OUTPUT = 'LONG'
SET FINAL_MALIGN3D = 1
CALL ROUTINE = 'very_fast' # prepare for extremely fast optimization
CALL ROUTINE = 'model' # do homology modelling
- How can I refine the model in successive steps?
There is a pre-defined routine 'select_atoms' which selects the atoms
to be moved during optimization. By default, the routine selects all
atoms, but you can redefine it to select any subset of atoms and then
only those atoms will be refined. They will ``feel'' the presence of other
atoms via all the static and possibly dynamic restraints that include
both selected and un-selected atoms.
For example, the script below would refine only atoms in residues 1 and 2
(file 'examples/tutorial-model/model-segment.top'). The difference
between this script and the one for loop modeling is that here
the selected regions are optimized with the default optimization protocol
and the default restraints, which generally include template-derived
restraints. In contrast, the loop modeling routine does not use
template-dependent restraints, but does a much more thorough optimization.
# Homology modelling by the MODELLER TOP routine 'model'.
# Demonstrates how to refine only a part of the model.
#
# You may want to use the more exhaustive "loop" modeling routines instead.
#
INCLUDE # Include the predefined TOP routines
SET OUTPUT_CONTROL = 1 1 1 1 0
SET ALNFILE = 'alignment.ali' # alignment filename
SET KNOWNS = '5fd1' # codes of the templates
SET SEQUENCE = '1fdx' # code of the target
SET ATOM_FILES_DIRECTORY = './:../atom_files' # directories for input atom files
SET STARTING_MODEL= 3 # index of the first model
SET ENDING_MODEL = 3 # index of the last model
# (determines how many models to calculate)
SET NONBONDED_SEL_ATOMS = 2 # selected atoms do not feel the neighbourhood
CALL ROUTINE = 'model' # do homology modelling
SUBROUTINE ROUTINE = 'select_atoms'
PICK_ATOMS SELECTION_SEGMENT='1:' '2:', SELECTION_SEARCH='segment', ;
PICK_ATOMS_SET=1, RES_TYPES='all', ATOM_TYPES='all', ;
SELECTION_FROM='all', SELECTION_STATUS='initialize'
RETURN
END_SUBROUTINE
- I want to model one or more loops very thoroughly (meaning spending
a lot of CPU time, not necessarily modeling more accurately).
Note that loops and insertions are already modeled by the default modeling
routine, so you do not have to do anything special to get a model for
the insertions. However, if you really want to focus on loops, you
can use the new loop modeling routine 'loop' (Section 3.3).
The selected regions are optimized independently
many times by a thorough molecular dynamics/simulated annealing procedure,
using sequence-dependent restraints only, no homology-derived restraints.
# Homology modelling by the MODELLER TOP routine 'model'.
# Demonstrates how to refine only a part of the model.
#
# This can be ran with run_clustor model-loop.top, too.
#
# The difference with model-segment is that the loop is
# refined on the basis of sequence alone, in the context
# of the rest of the structure.
INCLUDE # Include the predefined TOP routines
SET OUTPUT_CONTROL = 1 1 1 1 1
SET SEQUENCE = '1fdx' # code of the target
SET LOOP_MODEL = '1fdx.B99990001' # initial model of the target
SET ATOM_FILES_DIRECTORY = './:../atom_files' # directories for input atom files
# index of the first loop model:
SET LOOP_STARTING_MODEL = 20
# index of the last loop model:
SET LOOP_ENDING_MODEL = 23
SET LOOP_MD_LEVEL = 'refine_1' # the loop refinement method (1 fast / 3 slow)
CALL ROUTINE = 'loop'
# This routine picks model residues that need to be refined (necessary):
SUBROUTINE ROUTINE = 'select_loop_atoms'
# Uncomment if you also want to optimize the loop environment:
# SET SELECTION_SEARCH = 'SPHERE_SEGMENT', SPHERE_RADIUS = 6
# 4 residue insertion (1st loop):
PICK_ATOMS SELECTION_SEGMENT = '19:' '28:', SELECTION_STATUS = 'initialize'
# 2 residue insertion (2nd loop):
# PICK_ATOMS SELECTION_SEGMENT = '46:' '55:', SELECTION_STATUS = 'add'
RETURN
END_SUBROUTINE
# This routine adds any special restraints (optional):
#
# SUBROUTINE ROUTINE = 'special_restraints'
# MAKE_RESTRAINTS RESTRAINT_TYPE = 'ALPHA', RESIDUE_IDS = '46:' '55:'
# RETURN
# END_SUBROUTINE
- I want to build a model of a chimeric protein based on two
known structures. Alternatively, I want to build a multi-domain protein
model using templates corresponding only to the individual domains.
This can be accomplished using the standard modeling routine.
The alignment should be as follows when the chimera is a
combination of proteins A and B:
proteinA aaaaaaaaaaaaaaaaaaaaaaaaaaaa----------------------------------
proteinB ----------------------------bbbbbbbbbbbbbbbbbbbbbbbbbbbbbbbbbb
chimera aaaaaaaaaaaaaaaaaaaaaaaaaaaabbbbbbbbbbbbbbbbbbbbbbbbbbbbbbbbbb
In the PIR format the alignment file is:
>P1;proteinA
structureX:proteinA
aaaaaaaaaaaaaaaaaaaaaaaaaaaa----------------------------------*
>P1;proteinB
structureX:proteinB
----------------------------bbbbbbbbbbbbbbbbbbbbbbbbbbbbbbbbbb*
>P1;chimera
sequence:chimera
aaaaaaaaaaaaaaaaaaaaaaaaaaaabbbbbbbbbbbbbbbbbbbbbbbbbbbbbbbbbb*
If no additional information is available about the relative orientation
of the two domains the resulting model will probably have an incorrect
relative orientation of the two domains when the overlap between
A and B is non-existing or short. To obtain satisfactory
relative orientation of modeled domains in such cases, orient
the two template structures appropriately before the modeling.
- I don't want to use one region of a template for construction
of my model.
The easiest way to achieve this is to not align that region of the template
with the target sequence. If region 'bbbbbbbb' of the template should
not be used as a template for region 'eeeee' of the target sequence the
alignment should be like this:
template aaaaaaaaaaaaaaaaaaaaaaaa-----bbbbbbbbcccccccccccccccccccccccccccccc
target ddddddddddddddddddddddddeeeee--------ffffffffffffffffffffffffffffff
The effect of this alignment is that no homology-derived restraints will
be produced for region 'eeeee'.
- I want to define (additional) disulfide bonds in the target
sequence because no equivalent disulfide bonds exist in any of the
templates (in which case PATCH_SS_TEMPLATES cannot define them
automatically).
MODELLER can restrain disulfides in two ways: automatically
(PATCH_SS_TEMPLATES or
PATCH_SS_MODEL) and
manually (PATCH).
If there is an equivalent disulfide bridge in any
of the templates aligned with the target, the PATCH_SS_TEMPLATES
command will generate appropriate disulfide bond restraints without
any other input from the user. This command is run automatically by the
'model' script used for comparative modeling. The restraints include
bond, angle and dihedral angle restraints. The SG -- SG atom pair
also becomes an excluded atom pair that is not checked for an atom-atom
overlap. The 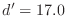 dihedral angle restraints will depend
on the conformation of the equivalent disulfides in the template
structure, as described in [Šali & Overington, 1994]. The command
PATCH_SS_MODEL is similar, except that the current structure of
MODEL, not templates, is used to guess the disulfide bonded
CYS - CYS pairs. dihedral angle restraints will depend
on the conformation of the equivalent disulfides in the template
structure, as described in [Šali & Overington, 1994]. The command
PATCH_SS_MODEL is similar, except that the current structure of
MODEL, not templates, is used to guess the disulfide bonded
CYS - CYS pairs.
Explicit manual restraints can be added by the PATCH
command relying on the PRES DISU patching residue in the CHARMM topology file. This command is used by the 'special_patches' routine
that is called automatically by the 'model' script. In comparative
modeling by 'model', the `manual' disulfides should be defined
in the 'special_patches' routine. The PATCH command will establish
the correct stereochemistry by relying on the CHARMM topology
file and parameters to restrain the disulfide bond.
It is better to use PATCH_SS_TEMPLATES than PATCH
where possible because the dihedral angles are restrained more
precisely by using the templates than the general rules of
stereochemistry.
Some CHARMM parameter files have a multiple dihedral entry for
the disulfide dihedral angle 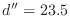 that consists of three individual
entries with periodicities of 1, 2 and 3. This is why you see three
feature restraints for a single disulfide in the output of the
ENERGY command. that consists of three individual
entries with periodicities of 1, 2 and 3. This is why you see three
feature restraints for a single disulfide in the output of the
ENERGY command.
# This is as usual:
INCLUDE
SET ALIGNMENT_FILE = 'align1.ali', KNOWNS='templ1', SEQUENCE='targ1'
CALL ROUTINE = 'model'
STOP
# Redefine the special_patches routine to include the additional disulfides
# (this routine is empty by default):
SUBROUTINE ROUTINE = 'special_patches'
# A disulfide between residues 1 and 85 in chain A:
PATCH RESIDUE_TYPE = 'DISU', RESIDUE_IDS = '1:A' '85:A'
# A disulfide between residues 41 and 45 in chain B:
PATCH RESIDUE_TYPE = 'DISU', RESIDUE_IDS = '41:B' '45:B'
RETURN
END_SUBROUTINE
- I want to explicitly force certain Pro residues to the
cis
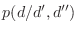 conformation. conformation.
MODELLER should usually be allowed to handle this automatically
via the omega dihedral angle restraints, which are calculated
by default.
# This is as usual:
INCLUDE
SET ALIGNMENT_FILE = 'align1.ali', KNOWNS='templ1', SEQUENCE='targ1'
CALL ROUTINE = 'model'
STOP
# Redefine the special_patches routine to force Pro to cis conformation:
# (this routine is empty by default):
SUBROUTINE ROUTINE = 'special_restraints'
CALL ROUTINE = 'cispeptide', ATOM_IDS1 = 'O:4' 'C:4' 'N:5' 'CA:5', ;
ATOM_IDS2 = 'CA:4' 'C:4' 'N:5' 'CA:5'
RETURN
END_SUBROUTINE
- How can I select/remove/add a set of restraints?
Restraints can be read from a file by READ_RESTRAINTS, calculated
by MAKE_RESTRAINTS, or added ``manually'' by ADD_RESTRAINT.
PICK_RESTRAINTS picks those restraints for objective function
calculation that restrain the selected atoms only, as specified
in the selected atoms set 1. Initially, all atoms are selected;
this can be changed by the PICK_ATOMS command.
MAKE_RESTRAINTS command for some restraint types (e.g., distance)
constructs restraints
of the selected type between the atoms in the selected atoms sets
2 and 3. Script 'scripts/__homcsr.top' contains examples
of the PICK_ATOMS command when generating restraints
by MAKE_RESTRAINTS. There are also commands
for adding and deleting single restraints, ADD_RESTRAINT
and DELETE_RESTRAINT, respectively. If you do CONDENSE_RESTRAINTS,
the unselected restraints will be deleted. This
is useful for getting rid of the unwanted restraints completely.
- I want to add my own restraints for optimization of the model.
You can read your restraints whenever the default restraints are read.
INCLUDE
SET ALIGNMENT_FILE = 'align1.ali', KNOWNS='templ1', SEQUENCE='targ1'
CALL ROUTINE = 'model'
STOP
# Redefine the rd_restraints routine:
SUBROUTINE ROUTINE = 'rd_restraints'
# This is the default homology-derived restraints:
READ_RESTRAINTS FILE = CSRFILE, ADD_RESTRAINTS = off
# This is two additional user provided files:
READ_RESTRAINTS FILE = 'my_rsrs1.rsr', ADD_RESTRAINTS = on
READ_RESTRAINTS FILE = 'my_rsrs2.rsr', ADD_RESTRAINTS = on
SET ADD_RESTRAINTS = off
RETURN
END_SUBROUTINE
- I want to add my own restraints to the file with the
automatically derived homology restraints, immediately after the
default calculation of the homology-derived restraints.
This is achieved by redefining the 'special_restraints' routine,
which is empty by default.
INCLUDE
SET ALIGNMENT_FILE = 'align1.ali', KNOWNS='templ1', SEQUENCE='targ1'
CALL ROUTINE = 'model'
# Redefine the special_restraints routine:
SUBROUTINE ROUTINE = 'special_restraints'
# Add some restraints from a file to existing homology-derived restraints:
READ_RESTRAINTS FILE = 'my_rsrs1.rsr', ADD_RESTRAINTS = on
# Restrain the specified CA-CA distance to 10 angstroms (st.dev.=0.1).
# Use a harmonic potential and X-Y distance group.
SET ATOM_IDS 'CA:35:A' 'CA:40:A'
ADD_RESTRAINT RESTRAINT_PARAMETERS = 3 1 1 27 2 2 0 10.0 0.1
SET ADD_RESTRAINTS = off
RETURN
END_SUBROUTINE
- I have my own restraints file to be used exclusively
for optimization by the default comparative modeling routine.
INCLUDE
SET ALIGNMENT_FILE = 'align1.ali', KNOWNS='templ1', SEQUENCE='targ1'
SET CSRFILE = 'targ1.rsr', CREATE_RESTRAINTS = 0
CALL ROUTINE = 'model'
- I have my own initial structure to be used for optimization
by the default comparative modeling routine.
INCLUDE
SET ALIGNMENT_FILE = 'align1.ali', KNOWNS='templ1', SEQUENCE='targ1'
# Specify the initial structure filename, and tell the program to
read the initial file, not construct it from the templates:
SET MODEL = 'targ1.ini', GENERATE_METHOD = 'read_xyz'
CALL ROUTINE = 'model'
- What are the different refinement levels really doing?
There are two different optimization approaches available within MODELLER:
variable target function method (VTFM) with conjugate gradients (CG)
[Šali & Blundell, 1993] and molecular dynamics (MD) with simulated
annealing (SA) [Šali & Blundell, 1993]. They can both be done to a different
degree (with more or less cycles of CG and MD, faster or slower schedule
for VTFM and SA). The exact details are best obtained from the scripts
themselves because a detailed description would probably be longer than
the scripts. For example, the QUANTA and INSIGHTII implementations
of MODELLER have these three levels of optimization:
no optimization (only copying coordinates from templates and
building the undefined atoms using internal geometry information from
the RTF entries); only VTFM with CG; also MD with SA. Most of the time
(70%) is spent on the MD&SA part. Our experience is that when MD&SA are
used, if there are violations in the best of the 10 models, they
probably come from an alignment error, not an optimizer failure
(if there are no insertions longer than approximately 15 residues).
- I want to change the default optimization schedule.
See file 'scripts/__defs.top' for the variables that could
be changed and for their possible values.
INCLUDE
SET ALIGNMENT_FILE = 'align1.ali', KNOWNS='templ1', SEQUENCE='targ1'
# Very thorough VTFM optimization:
SET LIBRARY_SCHEDULE = 1, MAX_VAR_ITERATIONS = 300
# Very thorough MD optimization:
SET MD_LEVEL = 'refine1'
# Repeat the whole cycle 3-times and do not stop unless obj.func. > 1E6
SET REPEAT_OPTIMIZATION = 3, MAX_MOLPDF = 1E6
CALL ROUTINE = 'model'
- I want to build an all hydrogen atom model with water molecules and
other non-protein atoms (atoms in the HETATM records in the PDB file).
INCLUDE
SET ALIGNMENT_FILE = 'align1.ali', KNOWNS='templ1', SEQUENCE='targ1'
SET TOPOLOGY_MODEL = 1, HYDROGEN_IO = on, HETATM_IO = on, WATER_IO = on
SET TOPLIB = $(LIB)/top.lib
SET PARLIB = $(LIB)/par.lib
CALL ROUTINE = 'model'
- How do I build a model with water molecules or residues that
do not have an entry in the topology and/or parameter files?
Water molecules are indicated by 'w' in the alignment file and the special
block residue ('BLK') that does not have entries in the residue topology
and parameter libraries is indicated by '.'
See Section 2.2.1 for information about block residues.
INCLUDE
SET ALIGNMENT_FILE = 'align1.ali', KNOWNS='templ1', SEQUENCE='targ1'
SET HETATM_IO = on, WATER_IO = on
CALL ROUTINE = 'model'
The alignment file:
>P1;templ1
structureX:templ1:1::10::
FAYVI/.wwww*
>P1;targ1
sequence:targ1:1::8::
-GWIV/.ww-w*
- How do I define my own residue types, such as D-amino acids,
special ligands, and unnatural amino-acids?
This is a painful area in all molecular modeling programs. However,
CHARMM and X-PLOR provide a reasonably straightforward solution
via the residue topology and parameter libraries. MODELLER uses
CHARMM topology and parameter library format and also extends the options
by allowing for a generic ``BLK'' residue type (Section 2.2.1).
This BLK residue type circumvents the need for editing any library files,
but it is not always possible to use it. Due to its conformational
rigidity, it is also not as accurate as a normal residue
type. In order to define a new residue type in the MODELLER
libraries, you have to follow the series of steps described below.
As an example, we will define the ALA residue without any hydrogen
atoms. You can add an entry to the MODELLER topology or parameter file;
you can also use your own topology or parameter files.
For more information, please see the CHARMM manual.
- Define the new residue entry in the residue topology file (RTF),
say 'top_heav.lib'.
RESI ALA 0.00000
ATOM N NH1 -0.29792
ATOM CA CT1 0.09563
ATOM CB CT3 -0.17115
ATOM C C 0.69672
ATOM O O -0.32328
BOND CB CA N CA O C C CA C +N
IMPR C CA +N O CA N C CB
IC -C N CA C 1.3551 126.4900 180.0000 114.4400 1.5390
IC N CA C +N 1.4592 114.4400 180.0000 116.8400 1.3558
IC +N CA *C O 1.3558 116.8400 180.0000 122.5200 1.2297
IC CA C +N +CA 1.5390 116.8400 180.0000 126.7700 1.4613
IC N C *CA CB 1.4592 114.4400 123.2300 111.0900 1.5461
IC N CA C O 1.4300 107.0000 0.0000 122.5200 1.2297
PATC FIRS NTER LAST CTER
You can obtain an initial approximation to this entry by defining the new
residue type using the residue type editor in QUANTA and then writing it
to a file.
The RESI record specifies the CHARMM residue name, which can be up to four
characters long and is usually the same as the PDB residue name (exceptions
are the potentially charged residues where the different charge states
correspond to different CHARMM residue types). The number gives the
total residue charge.
The ATOM records specify the IUPAC (i.e., PDB) atom names and the CHARMM atom
types for all the atoms in the residue.1.1 The number at the end of each ATOM record gives the partial
atomic charge.
The BOND records specify all the covalent bonds between the atoms in the
residue (e.g., there are bonds CB-CA, N-CA, O-C, etc.). In addition,
symbol '+' is used to indicate the bonds to the subsequent residue in the
chain (e.g., C - +N). The covalent angles and dihedral angles are calculated
automatically from the list of chemical bonds.
The IMPR records specify the improper dihedral angles, generally used
to restrain the planarity of various groups (e.g., peptide bonds and
sidechain rings). See also below.
The IC (internal coordinate) records are used for constructing
the initial Cartesian coordinates of a residue. An entry
specifies distances 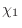 , angles , angles 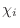 , and either dihedral angles
or improper dihedral angles , and either dihedral angles
or improper dihedral angles 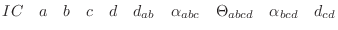 between atoms between atoms 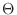 , , 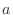 , , 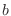 and and
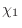 , given by their IUPAC names. The improper dihedral angle
is specified when the third atom, , given by their IUPAC names. The improper dihedral angle
is specified when the third atom, 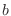 , is preceded by a star,
'*'. As before, the '-' and '+' pre-fixes for the atom names select
the corresponding atom from the preceding and subsequent residues,
respectively. The distances are in angstroms, angles in degrees.
The distinction between the dihedral angles and
improper dihedral angles is unfortunate since they are the
same mathematically, except that by convention when using the
equations, the order of the atoms for a dihedral angle is , is preceded by a star,
'*'. As before, the '-' and '+' pre-fixes for the atom names select
the corresponding atom from the preceding and subsequent residues,
respectively. The distances are in angstroms, angles in degrees.
The distinction between the dihedral angles and
improper dihedral angles is unfortunate since they are the
same mathematically, except that by convention when using the
equations, the order of the atoms for a dihedral angle is 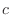 and for an improper dihedral angle it is
and for an improper dihedral angle it is 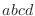 . .
The PATC record specifies the default patching residue type when the
current residue type is the first or the last residue in a chain.
- You have to make sure that all the CHARMM atom types of the
new residue type occur in the MASS records at the beginning
of the topology library: Add your entry at the end of the MASS list if
necessary. If you added any new CHARMM atom types, you also have to
add them to the radii libraries, 'modlib/radii.lib' and
'modlib/radii14.lib'. These libraries list the atomic radii for the
different topology models, for the long range and 1-4 non-bonded
soft-sphere terms, respectively. The full names of the files that are
used during calculation are given by the environment variables
$RADII_LIB and $RADII14_LIB.
- Optionally, you can add the residue entry to the library of
MODELLER topology models, 'modlib/models.lib'. The runtime
version of this library is specified by the environment variable
$MODELS_LIB. This library specifies which subsets of atoms
in the residue are used for each of the possible topologies.
Currently, there are 9 topologies selected by MODEL_TOPOLOGY
(3 is default):
| 1 |
ALLH |
all atoms |
| 2 |
POL |
polar hydrogens only |
| 3 |
HEAV |
non-hydrogen atoms only |
| 4 |
MCCB |
non-hydrogen mainchain (N, C, CA, O) and CB atoms |
| 5 |
MNCH |
non-hydrogen mainchain atoms only |
| 6 |
MCWO |
non-hydrogen mainchain atoms without carbonyl O |
| 7 |
CA |
CA atoms only |
| 8 |
MNSS |
non-hydrogen mainchain atoms and disulfide bonds |
| 9 |
CA3H |
reduced model with a small number of sidechain interaction centers |
The Ala entry is:
#
ALLH POLH HEAV MCCB MNCH MCWO CA MNSS CA3H
*
RESI ALA
ATOM NH1 NH1 NH1 NH1 NH1 NH1 #### NH1 ####
ATOM H HN #### #### #### #### #### #### ####
ATOM CT1 CT1 CT1 CT1 CT1 CT1 CT1 CT1 CAH
ATOM HB #### #### #### #### #### #### #### CH3E
ATOM CT3 CT3 CT3 CT3 #### #### #### #### ####
ATOM HA #### #### #### #### #### #### #### ####
ATOM HA #### #### #### #### #### #### #### ####
ATOM HA #### #### #### #### #### #### #### ####
ATOM C C C C C C #### C ####
ATOM O O O O O #### #### O ####
The residue entries in this library are separated by stars. The
'####' string indicates a missing atom. The atom names for the
present atoms are arbitrary. The order of the atoms must be the same
as in the CHARMM residue topology library. If a residue type
does not have an entry in this library, all atoms are used for
all topologies.
- You have to add the new residue type to the residue type library,
'modlib/restyp.lib'. The execution version of this file is
specified by the environment variable $RESTYP_LIB. For the
ALA residue,
1 | ALA | A | ALA | alanine
You would generally add the new residue type at the end of the
file. There are 5 fields in each line, separated by the '|' characters.
The first field is an integer index corresponding to the integer
residue type. The standard residue types have their indices smaller
than 24. These are also the indices corresponding to the residue-residue
substitution matrices. The second field contains the list of equivalent
PDB or IUPAC 3-character residue names, used in the PDB files. A list
rather than a single name is allowed because PDB can unfortunately use
different names for the same residue type (e.g., water can be HOH, WAT, etc.).
The third field gives a single character code for the residue type,
which is used in the alignment file. This does not have to be unique,
but if it is not unique you cannot use it in the alignment file.
Any ASCII character is fine, it does not have to be a letter. If you
run out of characters you can re-define the existing ones that you
do not need. The fourth field gives the four-character CHARMM residue
name, as specified in the RESI record of the topology library. The
last field contains an optional comment.
Every residue in the CHARMM topology file has to have an entry
in the $RESTYP_LIB library, but not every residue entry in the
$RESTYP_LIB library needs an entry in the residue topology file.
When you are adding a new residue type, you have to hope that
the maximal number of residue types is not over-reached. If it is,
a fatal error is reported at the beginning of the execution. To
solve this problem, you could delete some of the un-needed existing
residue types in the $RESTYP_LIB file, rather than re-compile the
program with larger array sizes. You can also read your own
residue type library by the READ_RESTYP_LIB command.
- In general, when you add a new residue type, you also add new
chemical bonds, angles, dihedral angles, improper dihedral angles,
and non-bonded interactions,
new in the sense that a unique combination of CHARMM atoms types
is involved whose interaction parameters are not yet specified in the
parameter library (see also Section 2.2.1).
In such a case, you will get a number of
warning and/or error messages when you generate the stereochemical
restraints by the MAKE_RESTRAINTS command. These messages
can sometimes be ignored because MODELLER will guess the
values for the missing parameters from the current Cartesian coordinates
of the model. When this is not accurate enough or if the necessary
coordinates are undefined
you have to specify the parameters explicitly in the parameter
library. Search for BOND, ANGL, DIHE, and IMPR sections in the
parameters library file and use the existing entries to guess your
new entries. Note that you can use dummy atom types 'X' to create
general dihedral (i.e., X A A X) and improper dihedral angle (i.e.,
A X X A) entries, where A stands for any of the real CHARMM
atom types.
For the dihedral angle cosine terms, the CHARMM convention for the phase
is different for 180
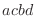 from MODELLER's (Eq. 5.56).
If you use non-bonded Lennard-Jones terms, you also have to add a NONB
entry for each new atom type. If you use the default soft-sphere
non-bonded restraints, you have already taken care of it by
adding the new atom types to the $RADII_LIB and $RADII_LIB
libraries. from MODELLER's (Eq. 5.56).
If you use non-bonded Lennard-Jones terms, you also have to add a NONB
entry for each new atom type. If you use the default soft-sphere
non-bonded restraints, you have already taken care of it by
adding the new atom types to the $RADII_LIB and $RADII_LIB
libraries.
- How do I define my own patching residue types?
This is even messier than defining a new residue type. As an example,
we will define the patching residue for establishing a disulfide
bond between two CYS residues.
PRES DISU -0.36 ! Patch for disulfides. Patch must be 1-CYS and 2-CYS.
ATOM 1CB CT2 -0.10 !
ATOM 1SG SM -0.08 ! 2SG--2CB--
ATOM 2SG SM -0.08 ! /
ATOM 2CB CT2 -0.10 ! -1CB--1SG
DELETE ATOM 1HG1
DELETE ATOM 2HG1
BOND 1SG 2SG
IC 1CA 1CB 1SG 2SG 0.0000 0.0000 180.0000 0.0000 0.0000
IC 1CB 1SG 2SG 2CB 0.0000 0.0000 90.0000 0.0000 0.0000
IC 1SG 2SG 2CB 2CA 0.0000 0.0000 180.0000 0.0000 0.0000
The PRES record specifies the CHARMM patching residue type (up to
four characters). As for the normal RESI residue types, patching
residue types also have to be defined in the residue type library,
'modlib/restyp.lib'.
The ATOM records have the same meaning as for the RESI residue types
described above. The extension is that the IUPAC atom names (listed
first) must be pre-fixed by the index of the residue that is patched.
In this example, there are two CYS residues that are patched, thus
the prefixes 1 and 2. When using the PATCH
command, the order of the patched residues specified by RESIDUE_IDS
must correspond to these indices (this is only important when the
patch is not symmetric, unlike the 'DISU' patch in this example).
DELETE records specify the atoms to be deleted, the two hydrogens bonded
to the two sulphurs in this case.
The BOND and IC (internal coordinate) records are the same as those for
the RESI residues, except that the atom names are prefixed with the
patched residue indices.
- Is it possible to restrain secondary structure in the
target sequence?
Yes. There are 'ALPHA', 'STRAND' and 'SHEET' restraint types
that the MAKE_RESTRAINTS command can generate. One specifies the
segment which is then restrained to the specified secondary structure
conformation. For example,
# This is as usual:
INCLUDE
SET ALIGNMENT_FILE = 'align1.ali', KNOWNS='templ1', SEQUENCE='targ1'
CALL ROUTINE = 'model'
STOP
# Redefine the special_restraints routine to include the secondary
# structure restraints (this routine is empty by default):
SUBROUTINE ROUTINE = 'special_restraints'
SET ADD_RESTRAINTS = on
# An alpha-helix:
MAKE_RESTRAINTS RESTRAINT_TYPE = 'alpha', RESIDUE_IDS = '20' '30'
# SET KEEP_DUPL_RESTR = 'new'
# Two strands:
MAKE_RESTRAINTS RESTRAINT_TYPE = 'STRAND', RESIDUE_IDS = '1' '6'
MAKE_RESTRAINTS RESTRAINT_TYPE = 'STRAND', RESIDUE_IDS = '9' '14'
# An anti-parallel sheet:
MAKE_RESTRAINTS RESTRAINT_TYPE = 'SHEET', ATOM_IDS = 'N:1' 'O:14', SHEET_H-BONDS = -5
RETURN
END_SUBROUTINE
- I want to patch the N-terminal or (C-terminal) residue (e.g.,
to model acetylation properly), but the PATCH command does not work.
This is probably because the N-terminus is patched by default with
the NTER patching residue (corresponding to -NH3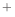 ) and a patched
residue must not be patched again. The solution is to turn the default
patching off by SET PATCH_DEFAULT = off before the
GENERATE_TOPOLOGY command is called. ) and a patched
residue must not be patched again. The solution is to turn the default
patching off by SET PATCH_DEFAULT = off before the
GENERATE_TOPOLOGY command is called.
- Is it possible to use templates with the coordinates for
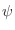 atoms only? atoms only?
Yes. You do not have to do anything special.
- How do I analyze the output log file?
First, check for the error messages by searching for string
'_E>''. These messages can only rarely be ignored. Next, check for the
warning messages by searching for string '_W>''. These messages
can almost always be ignored. If everything is OK so far, the most
important part of the log file is the output of
the ENERGY command for each model. This is where the violations of
restraints are listed. When there are too many too violated restraints,
more optimization or a different alignment is needed. What is too many
and too much? It depends on the restraint type and is best learned by
doing ENERGY on an X-ray structure or a good model to get a feel for it.
You may also want to look at the output of command CHECK_ALIGNMENT,
which should be self-explanatory. I usually ignore the other parts of the
log file.
- How do I prevent ``knots'' in the final models?
The best way to prevent knots is to start with a starting
structure that is as close to the desired final model as
possible. Other than that, the only solution at this point is to calculate
independently many models and hope that in some runs there won't
be knots. Knots usually occur when one or more neighboring long
insertions (i.e., longer than 15 residues) are modeled from
scratch. The reason is that an insertion is build from a randomized
distorted structure that is located approximately between the two
anchoring regions. Under such conditions, it is easy for the
optimizer to ``fall'' into a knot and then not be able to recover from it.
Sometimes knots result from an incorrect alignment, especially when
more than one template is used. When the alignment is correct,
knots are a result of optimization not being good enough.
However, making optimization more
thorough by increasing the CPU time would not be worth it on the
average as knots occur relatively infrequently. The excluded
volume restraints are already included in the
standard comparative modeling routine.
- What do I do when I get Syntax error at line 1: `(' unexpected message?
The executable is not recognized as such on your system. Make sure you FTP the file in the binary format.
Make sure the system version matches the self-descriptive name of the binary file.
Also it could be related to automatic processing of files by some Web browsers. Make sure you got a binary, not the file compressed by "compress" or "gzip" command. If you are not sure about the version of your system use the most generic executable which has been compiled for lower version of operating system.
- What is considered to be the minimum length of a sequence motif
necessary to derive meaningful constraints from the alignment to use in
modeling.. one, two, three, or more?
Usually more than that (dozens if you want just to detect reliable
similarity, and even more if you want a real model).
It is good to have at least 35-40% sequence identity to build a model.
Sometimes even 30% is OK.
- Does Modeller have a graphical interface (GUI) ?
No; Modeller is run from the command line, and uses a TOP script to direct it.
However, a graphical interface to Modeller is commercially available from
Accelrys, as part of Discovery Studio Modeling 1.1, at
http://www.accelrys.com/dstudio/ds_modeling/ds_modeler.html.
|